Bayesian Sampler Examples
Examples of running each sampler avaiable in 3ML.
Before, that, let’s discuss setting up configuration default sampler with default parameters. We can set in our configuration a default algorithm and default setup parameters for the samplers. This can ease fitting when we are doing exploratory data analysis.
With any of the samplers, you can pass keywords to access their setups. Read each pacakges documentation for more details.
[1]:
from threeML import *
from threeML.plugins.XYLike import XYLike
from packaging.version import Version
import numpy as np
import dynesty
from jupyterthemes import jtplot
%matplotlib inline
jtplot.style(context="talk", fscale=1, ticks=True, grid=False)
silence_warnings()
set_threeML_style()
05:50:37 WARNING The naima package is not available. Models that depend on it will not be functions.py:43 available
WARNING The GSL library or the pygsl wrapper cannot be loaded. Models that depend on it functions.py:65 will not be available.
WARNING The ebltable package is not available. Models that depend on it will not be absorption.py:33 available
[2]:
threeML_config.bayesian.default_sampler
[2]:
<Sampler.emcee: 'emcee'>
[3]:
threeML_config.bayesian.emcee_setup
[3]:
{'n_burnin': None, 'n_iterations': 500, 'n_walkers': 50, 'seed': 5123}
If you simply run bayes_analysis.sample() the default sampler and its default parameters will be used.
Let’s make some data to fit.
[4]:
sin = Sin(K=1, f=0.1)
sin.phi.fix = True
sin.K.prior = Log_uniform_prior(lower_bound=0.5, upper_bound=1.5)
sin.f.prior = Uniform_prior(lower_bound=0, upper_bound=0.5)
model = Model(PointSource("demo", 0, 0, spectral_shape=sin))
x = np.linspace(-2 * np.pi, 4 * np.pi, 20)
yerr = np.random.uniform(0.01, 0.2, 20)
xyl = XYLike.from_function("demo", sin, x, yerr)
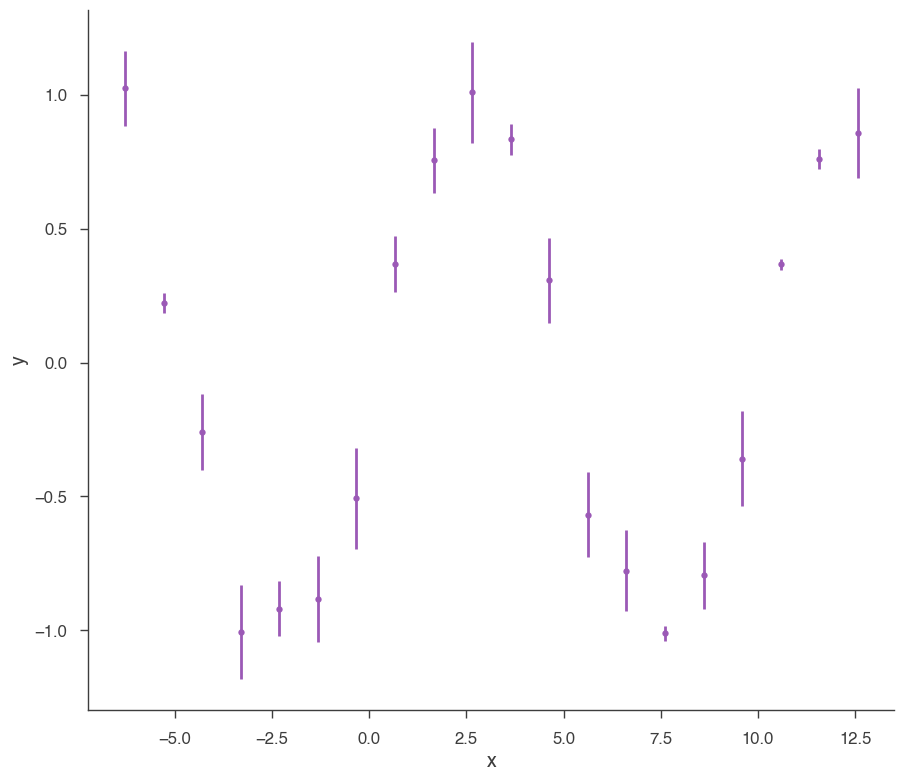
xyl.plot()
bayes_analysis = BayesianAnalysis(model, DataList(xyl))
emcee
[5]:
bayes_analysis.set_sampler("emcee")
bayes_analysis.sampler.setup(n_walkers=20, n_iterations=500)
bayes_analysis.sample()
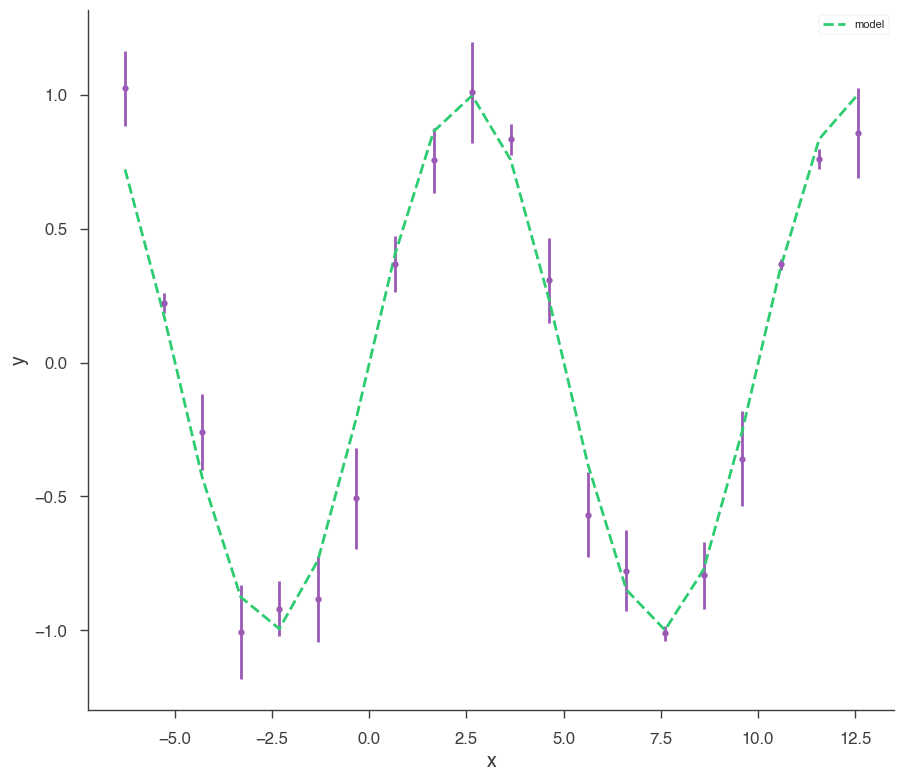
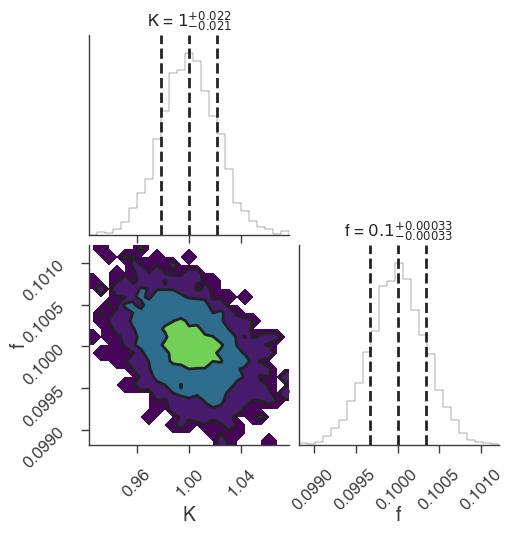
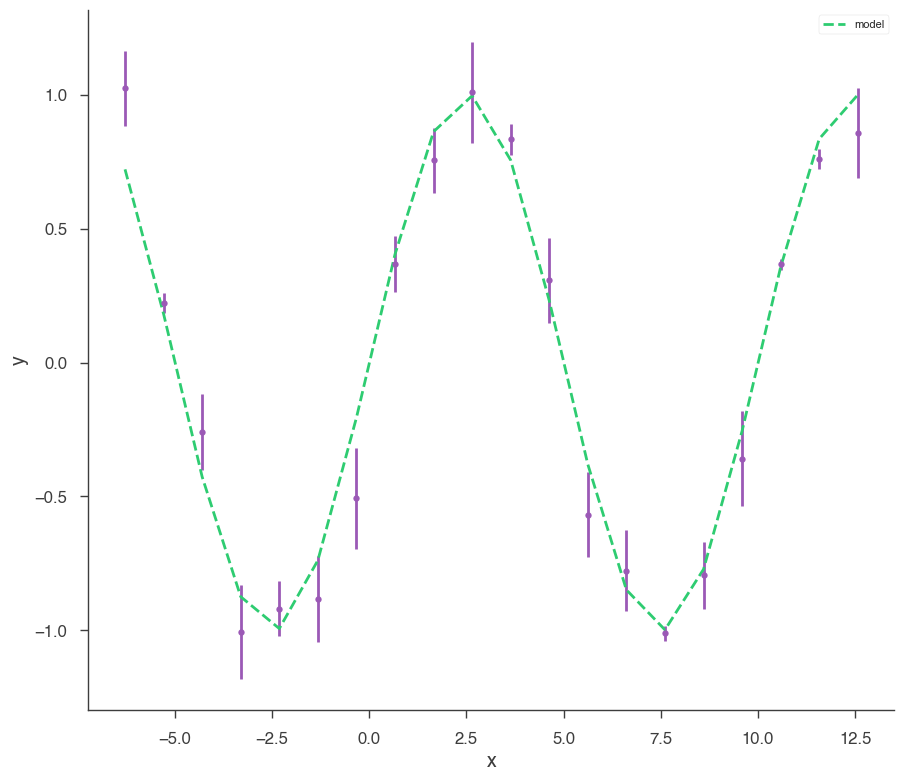
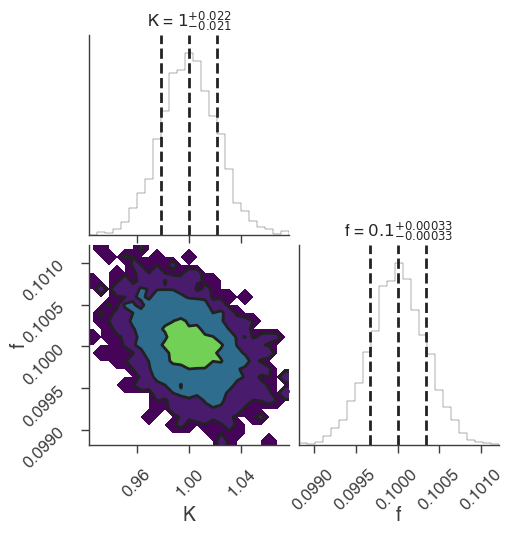
xyl.plot()
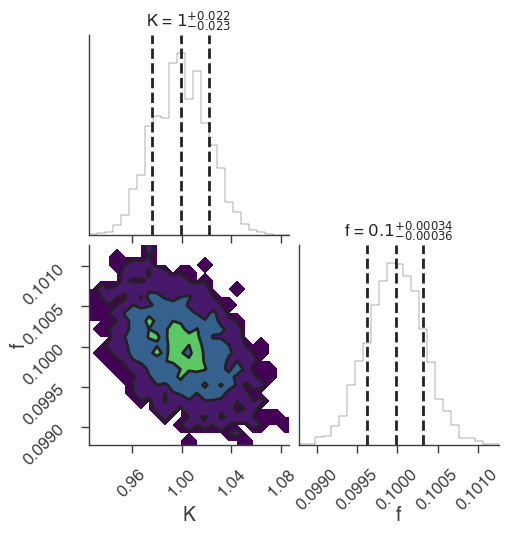
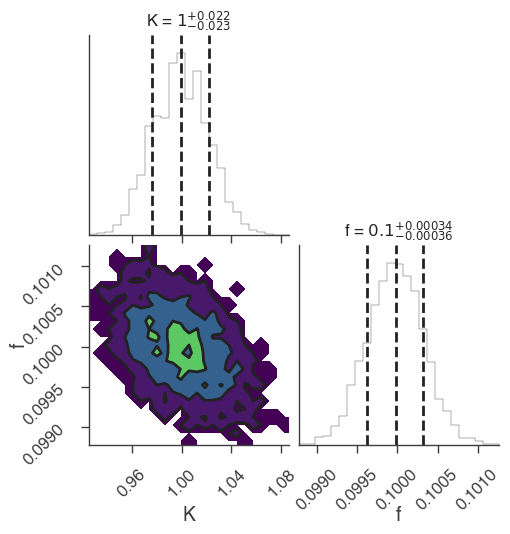
bayes_analysis.results.corner_plot()
INFO sampler set to emcee bayesian_analysis.py:186
05:50:41 INFO Mean acceptance fraction: 0.7278 emcee_sampler.py:145
INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.033 +/- 0.026 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.96 -0.08 +0.09) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.038918 |
| total | -9.038918 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.783718 |
| BIC | 24.069300 |
| DIC | 21.971510 |
| PDIC | 1.942418 |
[5]:
multinest
[6]:
bayes_analysis.set_sampler("multinest")
bayes_analysis.sampler.setup(n_live_points=400, resume=False, auto_clean=True)
bayes_analysis.sample()
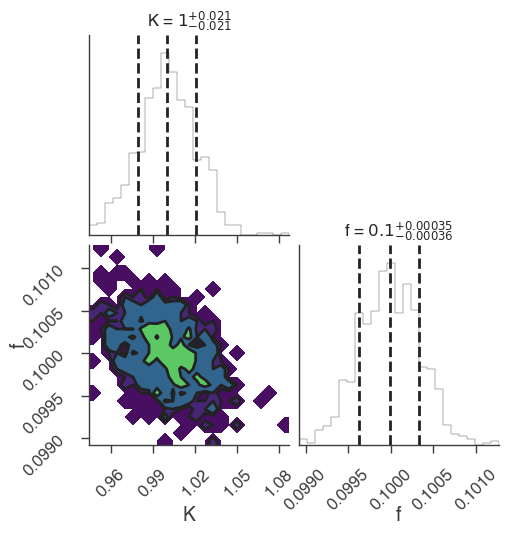
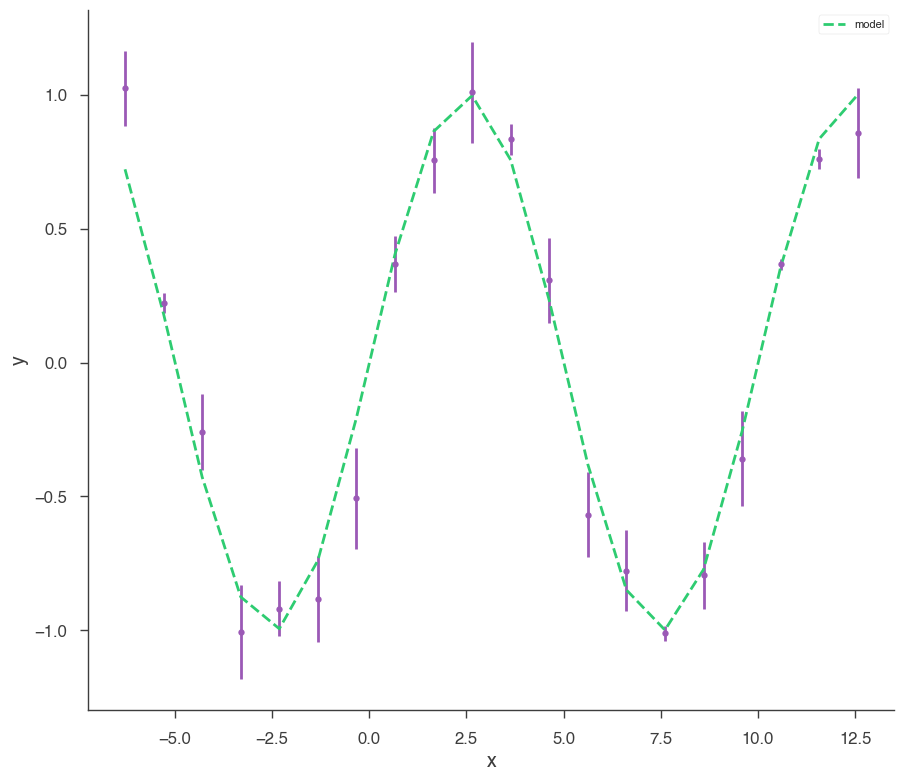
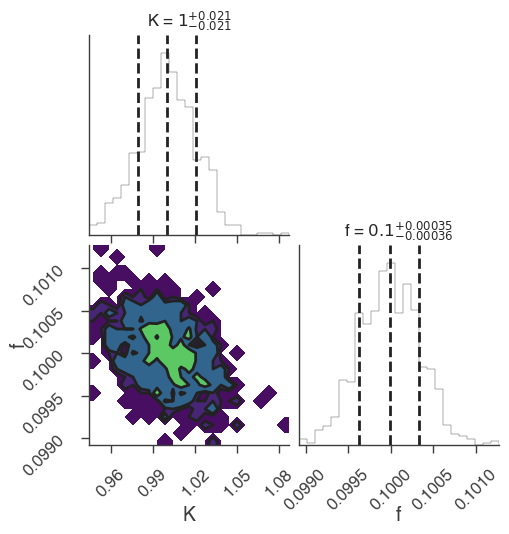
xyl.plot()
bayes_analysis.results.corner_plot()
05:50:42 INFO sampler set to multinest bayesian_analysis.py:186
*****************************************************
MultiNest v3.10
Copyright Farhan Feroz & Mike Hobson
Release Jul 2015
no. of live points = 400
dimensionality = 2
*****************************************************
MultiNest Warning!
Parameter 1 of mode 3 is converging towards the edge of the prior.
MultiNest Warning!
Parameter 1 of mode 3 is converging towards the edge of the prior.
analysing data from chains/fit-.txt ln(ev)= -17.055684536516001 +/- 0.13268055531958889
Total Likelihood Evaluations: 6008
Sampling finished. Exiting MultiNest
05:50:43 INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.032 -0.027 +0.028 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.96 +/- 0.08) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.039276 |
| total | -9.039276 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.784435 |
| BIC | 24.070017 |
| DIC | 22.119436 |
| PDIC | 2.020380 |
| log(Z) | -7.407190 |
INFO deleting the chain directory chains multinest_sampler.py:234
[6]:
dynesty
[7]:
bayes_analysis.set_sampler("dynesty_nested")
bayes_analysis.sampler.setup(nlive=400)
bayes_analysis.sample()
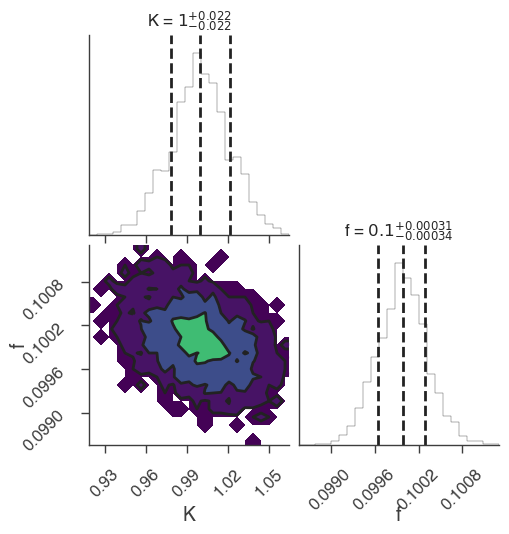
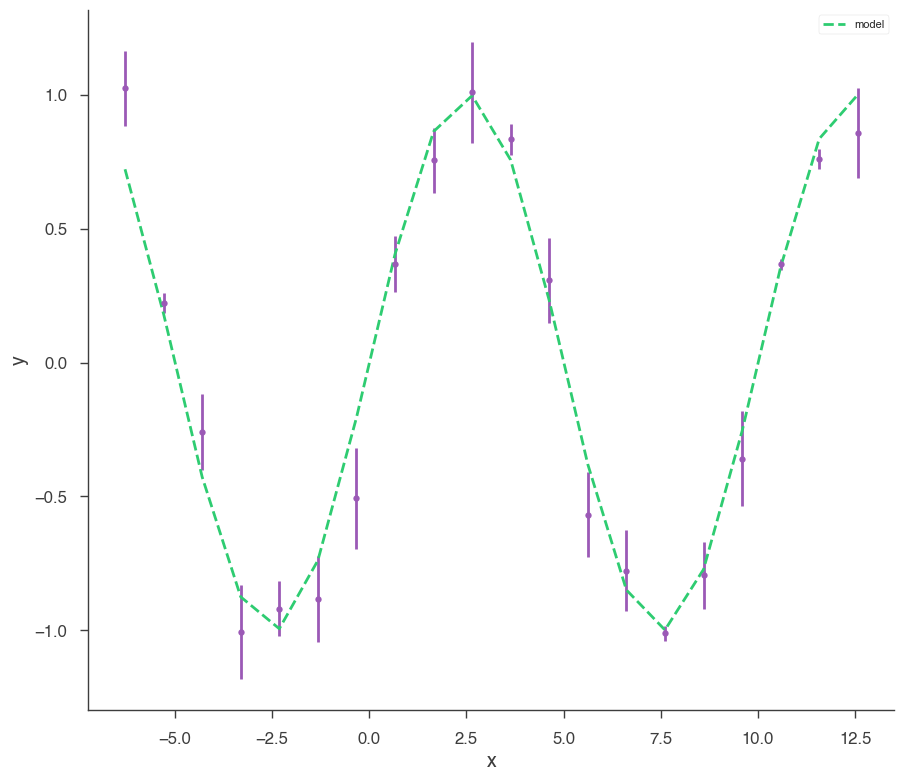
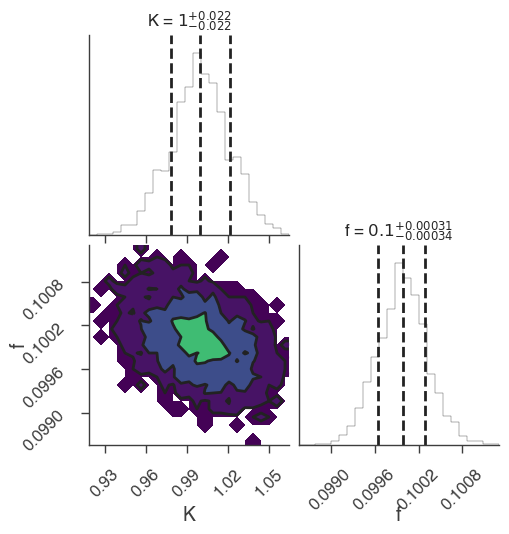
xyl.plot()
bayes_analysis.results.corner_plot()
INFO sampler set to dynesty_nested bayesian_analysis.py:186
3765it [00:03, 1153.31it/s, +400 | bound: 10 | nc: 1 | ncall: 18053 | eff(%): 23.594 | loglstar: -inf < -9.007 < inf | logz: -17.371 +/- 0.136 | dlogz: 0.001 > 0.409]
05:50:46 INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.033 -0.026 +0.027 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.95 -0.09 +0.08) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.039186 |
| total | -9.039186 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.784254 |
| BIC | 24.069836 |
| DIC | 22.086992 |
| PDIC | 2.004012 |
| log(Z) | -7.544179 |
[7]:
[8]:
bayes_analysis.set_sampler("dynesty_dynamic")
bayes_analysis.sampler.setup()
if Version(dynesty.__version__) >= Version("3.0.0"):
bayes_analysis.sample(n_effective=None)
else:
bayes_analysis.sample(
stop_function=dynesty.utils.old_stopping_function, n_effective=None
)
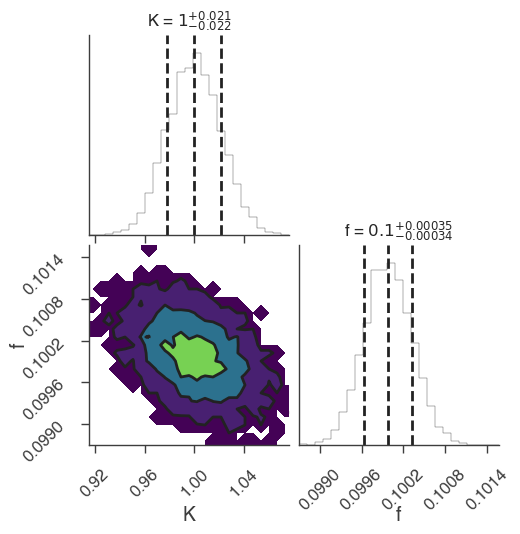
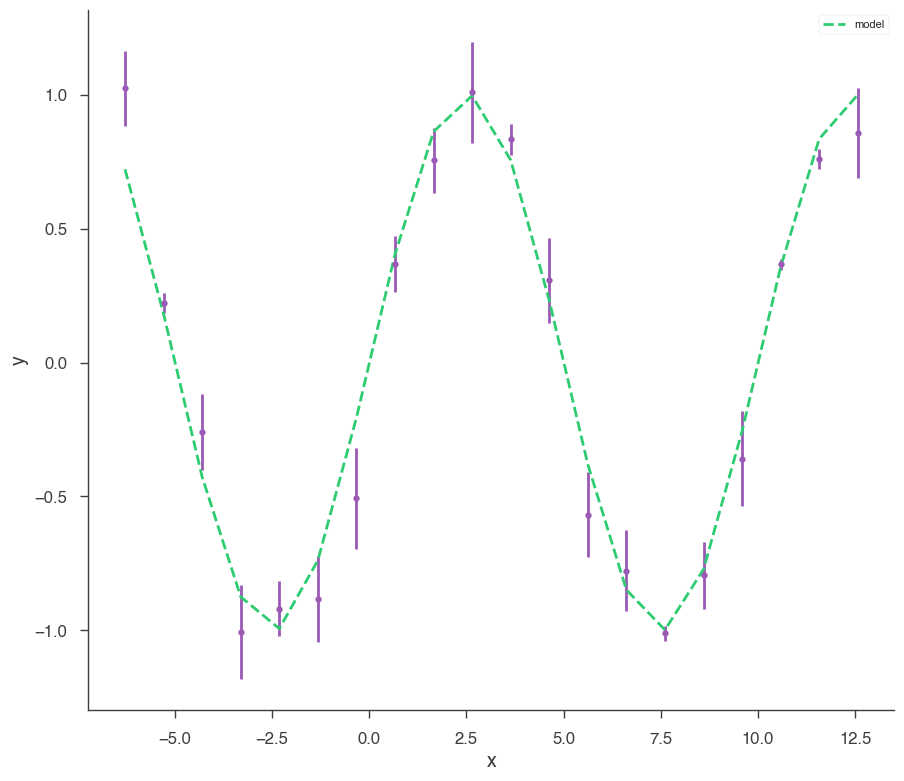
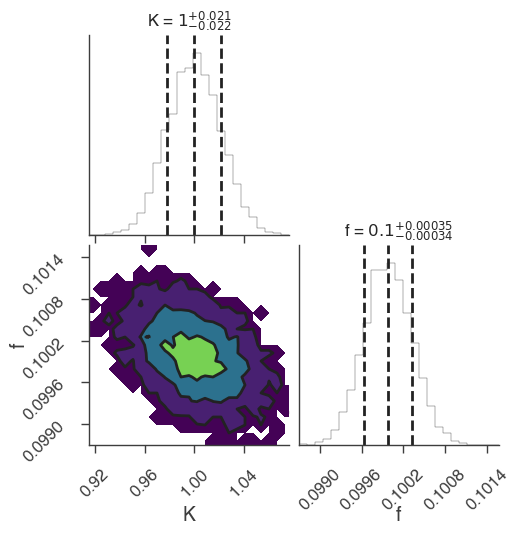
xyl.plot()
bayes_analysis.results.corner_plot()
05:50:47 INFO sampler set to dynesty_dynamic bayesian_analysis.py:186
15823it [00:13, 1169.82it/s, batch: 8 | bound: 4 | nc: 1 | ncall: 37322 | eff(%): 42.334 | loglstar: -13.988 < -9.007 < -9.356 | logz: -17.398 +/- 0.070 | stop: 0.898]
05:51:01 INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.033 +/- 0.027 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.96 -0.09 +0.08) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.038935 |
| total | -9.038935 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.783752 |
| BIC | 24.069334 |
| DIC | 22.071711 |
| PDIC | 1.996451 |
| log(Z) | -7.543777 |
[8]:
zeus
[9]:
bayes_analysis.set_sampler("zeus")
bayes_analysis.sampler.setup(n_walkers=20, n_iterations=500)
bayes_analysis.sample()
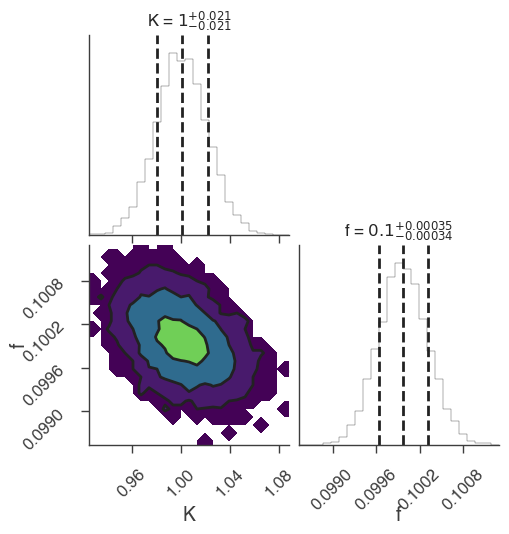
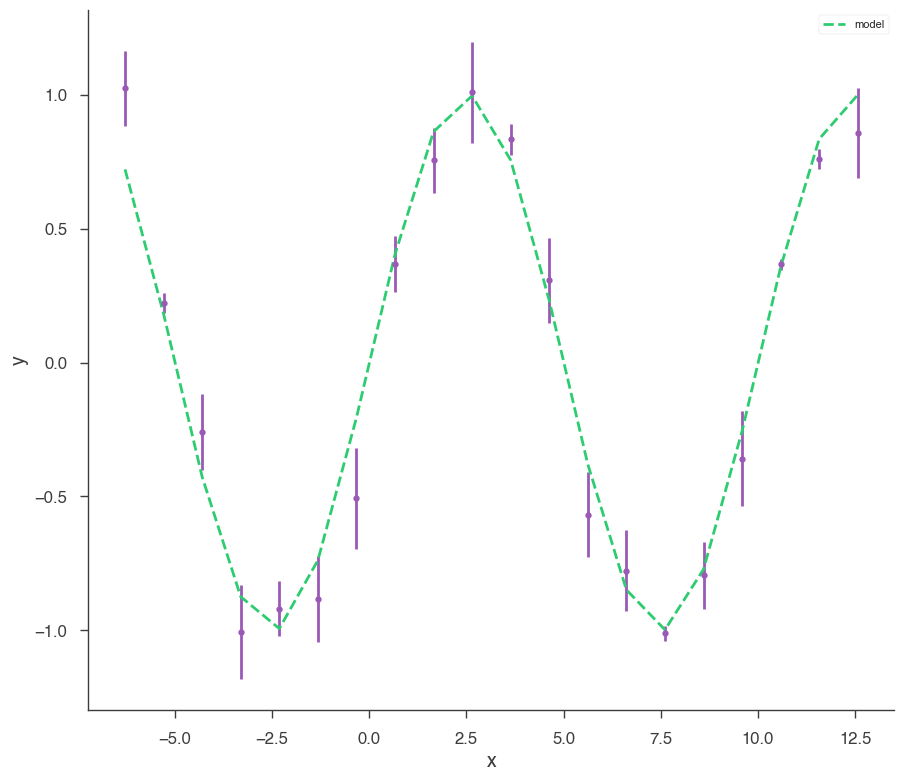
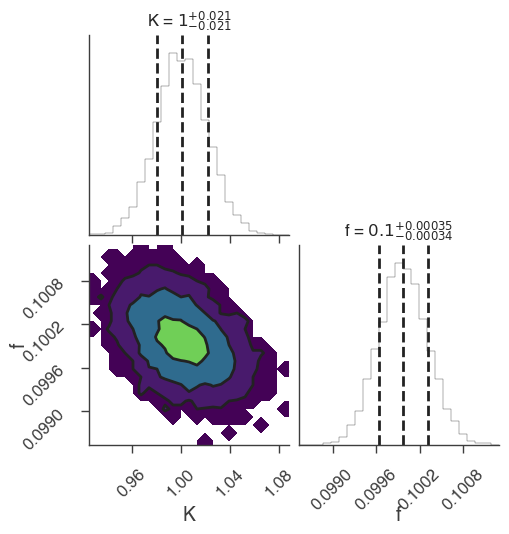
xyl.plot()
bayes_analysis.results.corner_plot()
INFO sampler set to zeus bayesian_analysis.py:186
The run method has been deprecated and it will be removed. Please use the new run_mcmc method.
Initialising ensemble of 20 walkers...
Sampling progress : 100%|██████████| 625/625 [00:03<00:00, 170.26it/s]
05:51:05 INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Summary
-------
Number of Generations: 625
Number of Parameters: 2
Number of Walkers: 20
Number of Tuning Generations: 38
Scale Factor: 1.249892
Mean Integrated Autocorrelation Time: 3.04
Effective Sample Size: 4112.68
Number of Log Probability Evaluations: 64917
Effective Samples per Log Probability Evaluation: 0.063353
None
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.033 -0.027 +0.025 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.96 -0.09 +0.08) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.038916 |
| total | -9.038916 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.783714 |
| BIC | 24.069296 |
| DIC | 22.003078 |
| PDIC | 1.961844 |
[9]:
ultranest
[10]:
bayes_analysis.set_sampler("ultranest")
bayes_analysis.sampler.setup(
min_num_live_points=400, frac_remain=0.5, use_mlfriends=False
)
bayes_analysis.sample()
xyl.plot()
bayes_analysis.results.corner_plot()
INFO sampler set to ultranest bayesian_analysis.py:186
[ultranest] Sampling 400 live points from prior ...
[ultranest] Explored until L=-9
[ultranest] Likelihood function evaluations: 5391
[ultranest] logZ = -17.19 +- 0.1132
[ultranest] Effective samples strategy satisfied (ESS = 974.8, need >400)
[ultranest] Posterior uncertainty strategy is satisfied (KL: 0.47+-0.09 nat, need <0.50 nat)
[ultranest] Evidency uncertainty strategy is satisfied (dlogz=0.42, need <0.5)
[ultranest] logZ error budget: single: 0.13 bs:0.11 tail:0.41 total:0.42 required:<0.50
[ultranest] done iterating.
05:51:08 INFO fit restored to maximum of posterior sampler_base.py:168
INFO fit restored to maximum of posterior sampler_base.py:168
Maximum a posteriori probability (MAP) point:
| result | unit | |
|---|---|---|
| parameter | ||
| demo.spectrum.main.Sin.K | 1.033 +/- 0.026 | 1 / (keV s cm2) |
| demo.spectrum.main.Sin.f | (9.95 -0.09 +0.08) x 10^-2 | rad / keV |
Values of -log(posterior) at the minimum:
| -log(posterior) | |
|---|---|
| demo | -9.039284 |
| total | -9.039284 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 22.784450 |
| BIC | 24.070032 |
| DIC | 22.120489 |
| PDIC | 2.020420 |
| log(Z) | -7.479574 |
[10]: