Joint analysis of GRB 130427A with GBM, LLE and LAT data
[1]:
import warnings
warnings.simplefilter("ignore")
from threeML import *
from threeML.utils.data_download.Fermi_LAT.download_LAT_data import LAT_dataset
import matplotlib.pyplot as plt
import numpy as np
import astropy.units as u
from astropy.io import fits
%matplotlib inline
06:28:02 WARNING The naima package is not available. Models that depend on it will not be functions.py:43 available
WARNING The GSL library or the pygsl wrapper cannot be loaded. Models that depend on it functions.py:65 will not be available.
WARNING The ebltable package is not available. Models that depend on it will not be absorption.py:33 available
[2]:
gbm_catalog = FermiGBMBurstCatalog()
source_name = "GRB130427324"
gbm_catalog.query_sources(source_name)
grb_info = gbm_catalog.get_detector_information()[source_name]
grb_info
06:28:04 INFO The cache for fermigbrst does not yet exist. We will try to get_heasarc_table_as_pandas.py:63 build it
INFO Building cache for fermigbrst get_heasarc_table_as_pandas.py:103
[2]:
{'source': {'fluence': '4.096000-142.338000', 'peak': '8.192000-9.216000'},
'background': {'pre': '-285.420000--41.670000',
'post': '381.250000-500.000000',
'full': '-285.420000--41.670000,381.250000-500.000000'},
'trigger': 'bn130427324',
'detectors': array(['n6', 'n9', 'na', 'b1'], dtype='<U2'),
'best fit model': {'fluence': 'band', 'peak': 'band'},
'ra': 173.136,
'dec': 27.7129}
[3]:
trigger_name = "bn130427324"
met = 388741629.420
ra = 173.136
dec = 27.7129
tstart = 0
tstop = 100
gbm_detectors = ["n6", "n9", "b1"]
roi = 10
zmax = 100.0
thetamax = 180.0
irf = "p8_transient010e"
Prepare GBM data
[4]:
dload = download_GBM_trigger_data(trigger_name, detectors=gbm_detectors)
[5]:
bkg_selection = ["-285--41", "310-355"]
gbm_plugins = []
time_series = {}
for det in gbm_detectors:
# Use CSPEC data to fit the background using the background selections.
# We use CSPEC because it has a longer duration for fitting the background.
ts_cspec = TimeSeriesBuilder.from_gbm_cspec_or_ctime(
det, cspec_or_ctime_file=dload[det]["cspec"], rsp_file=dload[det]["rsp"]
)
ts_cspec.set_background_interval(*bkg_selection)
# The background is saved to an HDF5 file that stores the polynomial coefficients and selections
ts_cspec.save_background(f"{det}_bkg.h5", overwrite=True)
# We use TTE data for the actual spectral analysis
ts_tte = TimeSeriesBuilder.from_gbm_tte(
det,
tte_file=dload[det]["tte"],
rsp_file=dload[det]["rsp"],
restore_background=f"{det}_bkg.h5",
)
time_series[det] = ts_tte
# The source selection from the catalog is set
ts_tte.set_active_time_interval("%f-%f" % (tstart, tstop))
# The plugin for the time integrated analysis is created for each detector
fluence_plugin = ts_tte.to_spectrumlike()
# GBM channel selections for spectral analysis
if det.startswith("b"):
fluence_plugin.set_active_measurements("900-30000")
else:
fluence_plugin.set_active_measurements("40-900")
fluence_plugin.rebin_on_background(1.0)
gbm_plugins.append(fluence_plugin)
06:29:17 WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING RuntimeWarning: invalid value encountered in divide
WARNING RuntimeWarning: invalid value encountered in divide
06:29:18 WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
06:29:19 INFO Auto-determined polynomial order: 1 binned_spectrum_series.py:356
06:29:26 INFO None 1-order polynomial fit with the mle method time_series.py:426
INFO Saved fitted background to n6_bkg.h5 time_series.py:972
INFO Saved background to n6_bkg.h5 time_series_builder.py:430
WARNING The TTE file gbm_data.py:38 /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/glg_tte_n6_bn130427324 _v00.fit.gz contains duplicate time tags and is thus invalid. Contact the FSSC
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
INFO Successfully restored fit from n6_bkg.h5 time_series_builder.py:166
06:29:27 INFO Interval set to 0.0-100.0 for n6 time_series_builder.py:274
INFO Auto-probed noise models: SpectrumLike.py:486
INFO - observation: poisson SpectrumLike.py:487
INFO - background: gaussian SpectrumLike.py:488
INFO Range 40-900 translates to channels 26-123 SpectrumLike.py:1237
06:29:31 INFO Now using 98 bins SpectrumLike.py:1706
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING RuntimeWarning: invalid value encountered in divide
WARNING RuntimeWarning: invalid value encountered in divide
06:29:32 WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
06:29:33 INFO Auto-determined polynomial order: 2 binned_spectrum_series.py:356
06:29:40 INFO None 2-order polynomial fit with the mle method time_series.py:426
INFO Saved fitted background to n9_bkg.h5 time_series.py:972
INFO Saved background to n9_bkg.h5 time_series_builder.py:430
WARNING The TTE file gbm_data.py:38 /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/glg_tte_n9_bn130427324 _v00.fit.gz contains duplicate time tags and is thus invalid. Contact the FSSC
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
INFO Successfully restored fit from n9_bkg.h5 time_series_builder.py:166
WARNING Minimum MC energy (5.0) is larger than minimum EBOUNDS energy response.py:133 (4.410999774932861)
INFO Interval set to 0.0-100.0 for n9 time_series_builder.py:274
INFO Auto-probed noise models: SpectrumLike.py:486
INFO - observation: poisson SpectrumLike.py:487
INFO - background: gaussian SpectrumLike.py:488
INFO Range 40-900 translates to channels 27-124 SpectrumLike.py:1237
INFO Now using 98 bins SpectrumLike.py:1706
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING RuntimeWarning: invalid value encountered in divide
WARNING RuntimeWarning: invalid value encountered in divide
06:29:42 WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
INFO Auto-determined polynomial order: 2 binned_spectrum_series.py:356
06:29:49 INFO None 2-order polynomial fit with the mle method time_series.py:426
INFO Saved fitted background to b1_bkg.h5 time_series.py:972
INFO Saved background to b1_bkg.h5 time_series_builder.py:430
WARNING The TTE file gbm_data.py:38 /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/glg_tte_b1_bn130427324 _v00.fit.gz contains duplicate time tags and is thus invalid. Contact the FSSC
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 2) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 3) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 4) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 5) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 6) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
06:29:50 WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 7) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 8) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 9) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', response.py:450 10) not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX' 'SPECRESP MATRIX'
WARNING No TLMIN keyword found. This DRM does not follow OGIP standards. Assuming response.py:576 TLMIN=1
INFO Successfully restored fit from b1_bkg.h5 time_series_builder.py:166
INFO Interval set to 0.0-100.0 for b1 time_series_builder.py:274
INFO Auto-probed noise models: SpectrumLike.py:486
INFO - observation: poisson SpectrumLike.py:487
INFO - background: gaussian SpectrumLike.py:488
INFO Range 900-30000 translates to channels 20-119 SpectrumLike.py:1237
INFO Now using 100 bins SpectrumLike.py:1706
[6]:
time_series["n6"].view_lightcurve(-10, 100)
time_series["n9"].view_lightcurve(-10, 100)
time_series["b1"].view_lightcurve(-10, 100)
[6]:
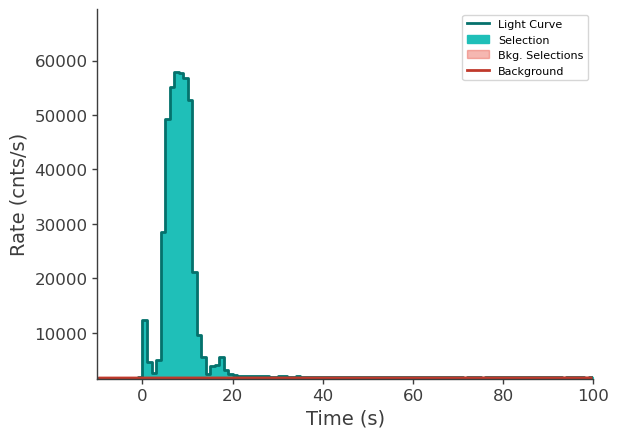
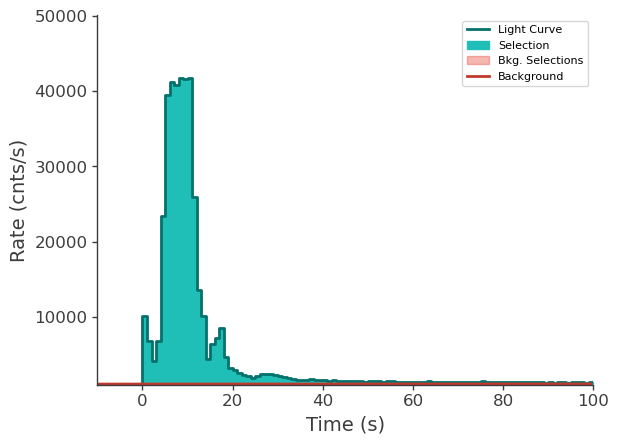
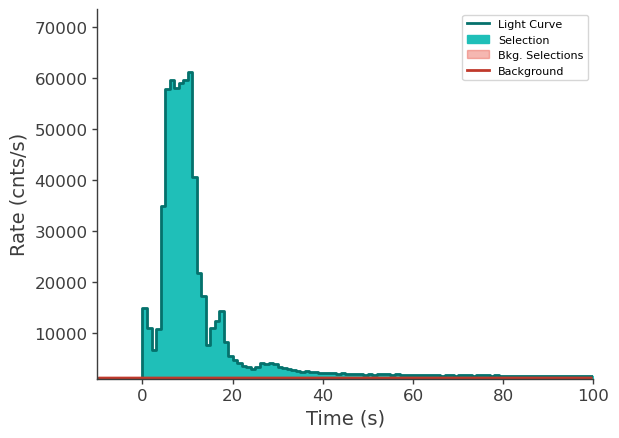
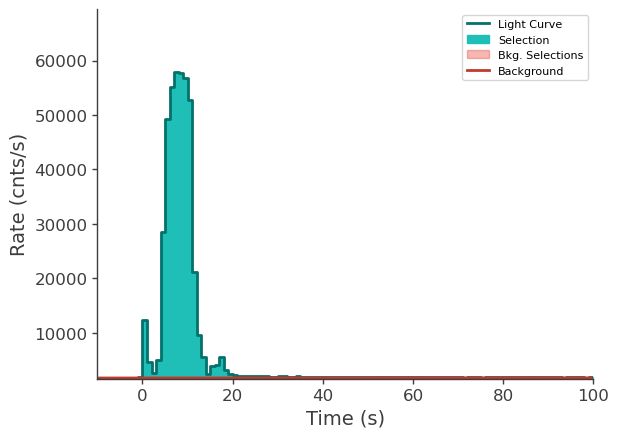
Prepare LLE data
[7]:
lle_dload = download_LLE_trigger_data(trigger_name)
[8]:
bkg_selection = ["-285--41", "400-500"]
emin, emax = 30 * u.MeV, 100 * u.MeV
lle_time_series = TimeSeriesBuilder.from_lat_lle(
"lat_lle",
lle_file=lle_dload["lle"],
ft2_file=lle_dload["ft2"],
rsp_file=lle_dload["rsp"],
)
lle_time_series.set_background_interval(*bkg_selection)
lle_time_series.save_background("lle_bkg.h5", overwrite=True)
lle_time_series.set_active_time_interval("%d-%d" % (tstart, tstop))
lle_plugin = lle_time_series.to_spectrumlike()
lle_plugin.set_active_measurements(
"%d-%d" % (emin.to("keV").value, emax.to("keV").value)
)
lle_plugin.use_effective_area_correction(0.8, 1.2)
06:29:52 WARNING The default choice for MATRIX extension failed:KeyError("Extension ('MATRIX', 1) response.py:450 not found.")available: None 'EBOUNDS' 'SPECRESP MATRIX'
INFO Auto-determined polynomial order: 2 event_list.py:505
06:29:55 INFO None 2-order polynomial fit with the mle method time_series.py:426
INFO Saved fitted background to lle_bkg.h5 time_series.py:972
INFO Saved background to lle_bkg.h5 time_series_builder.py:430
INFO Interval set to 0.0-100.0 for lat_lle time_series_builder.py:274
INFO Auto-probed noise models: SpectrumLike.py:486
INFO - observation: poisson SpectrumLike.py:487
INFO - background: gaussian SpectrumLike.py:488
INFO Range 30000-100000 translates to channels 5-12 SpectrumLike.py:1237
INFO lat_lle is using effective area correction (between 0.8 and 1.2) SpectrumLike.py:2224
[9]:
lle_time_series.view_lightcurve(-100, 500)
[9]:
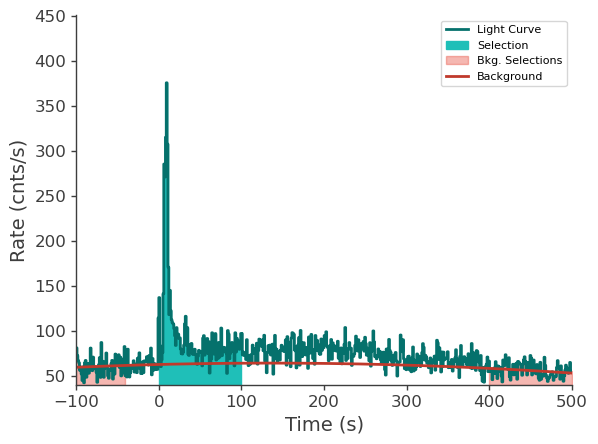
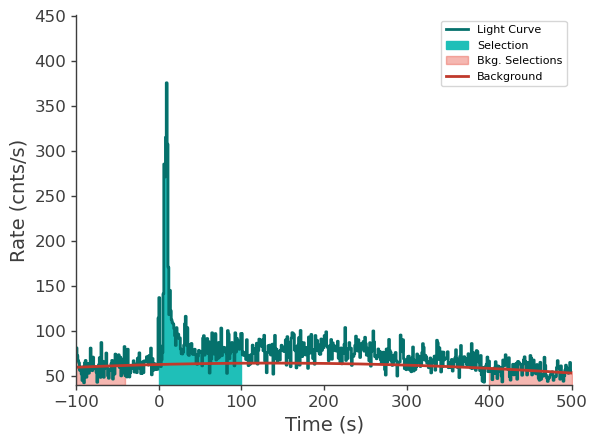
Prepare LAT data
[10]:
LATdataset = LAT_dataset()
LATdataset.make_LAT_dataset(
ra=ra,
dec=dec,
radius=12,
trigger_time=met,
tstart=0,
tstop=1000,
data_type="Extended",
destination_directory=".",
Emin=100.0, # 100 MeV
Emax=100000.0, # 100 GeV
)
INFO Query parameters: download_LAT_data.py:255
INFO coordfield = 173.1360,27.7129 download_LAT_data.py:258
INFO coordsystem = J2000 download_LAT_data.py:258
INFO shapefield = 12 download_LAT_data.py:258
INFO timefield = 388741629.42,388742629.42 download_LAT_data.py:258
INFO timetype = MET download_LAT_data.py:258
INFO energyfield = 100.000,100000.000 download_LAT_data.py:258
INFO photonOrExtendedOrNone = Extended download_LAT_data.py:258
INFO destination = query download_LAT_data.py:258
INFO spacecraft = checked download_LAT_data.py:258
INFO Query ID: ec3bd58c9484a4abd12bcf252f186da3 download_LAT_data.py:262
06:29:56 INFO Estimated complete time for your query: 16 seconds download_LAT_data.py:395
INFO If this download fails, you can find your data at download_LAT_data.py:404 https://fermi.gsfc.nasa.gov/cgi-bin/ssc/LAT/QueryResults.cgi?id=L2604300 229574CC8A8EB97 (when ready)
INFO We are gonna wait another 5.0s and retry download_LAT_data.py:461
06:30:02 INFO We are gonna wait another 5.0s and retry download_LAT_data.py:461
06:30:07 INFO Downloading FT1 and FT2 files... download_LAT_data.py:474
06:30:08 INFO ['https://fermi.gsfc.nasa.gov/FTP/fermi/data/lat/queries/L2604300229574 download_LAT_data.py:477 CC8A8EB97_EV00.fits', 'https://fermi.gsfc.nasa.gov/FTP/fermi/data/lat/queries/L2604300229574CC 8A8EB97_SC00.fits']
WARNING Only one FT1 file provided. Skipping the merge... download_LAT_data.py:87
Writing ./bn130427324/gll_cspec_tr_bn130427324_v00.rsp...
time -p gtselect infile=./bn130427324/gll_ft1_tr_bn130427324_v00.fit outfile=__temp_ft1.fits ra=173.136 dec=27.7129 rad=15.0 tmin=388741628.42 tmax=388742630.42 emin=10.0 emax=1000000.0 zmin=0.0 zmax=110.0 evclass="INDEF" evtype="INDEF" convtype=-1 phasemin=0.0 phasemax=1.0 evtable="EVENTS" chatter=2 clobber=yes debug=no gui=no mode="ql"
Done.
real 0.05
user 0.05
sys 0.00
Writing ./bn130427324/gll_cspec_tr_bn130427324_v00.pha...
* Get energy binning from the response matrix...
done.
* Run gtbindef and gtbin and bin in energy and time...
time -p gtbindef bintype="E" binfile=__ebins.txt outfile=__energyBins.fits energyunits="keV" chatter=2 clobber=yes debug=no gui=no mode="ql"
This is gtbindef version HEAD
real 0.00
user 0.00
sys 0.00
time -p gtbin evfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/__temp_ft1.fits scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit outfile=__gtllebin__pha2.pha algorithm="PHA2" ebinalg="FILE" emin=30.0 emax=200000.0 enumbins=0 denergy=0.0 ebinfile=__energyBins.fits tbinalg="LIN" tstart=388741629.42 tstop=388742629.42 dtime=4.096 tbinfile=NONE snratio=0.0 lcemin=0.0 lcemax=0.0 nxpix=0 nypix=0 binsz=0.0 coordsys="CEL" xref=0.0 yref=0.0 axisrot=0.0 rafield="RA" decfield="DEC" proj="AIT" hpx_ordering_scheme="RING" hpx_order=3 hpx_ebin=yes hpx_region="" evtable="EVENTS" sctable="SC_DATA" efield="ENERGY" tfield="TIME" chatter=1 clobber=yes debug=no gui=no mode="ql"
This is gtbin version HEAD
real 1.91
user 1.83
sys 0.08
done.
* Transform gtbin output in CSPEC format...
done.
* Updating keywords in the headers of the CSPEC file...
done.
gtllebin done!
[11]:
LATdataset.extract_events(roi, zmax, irf)
with fits.open(LATdataset.filt_file) as event_file:
lat_events = event_file["EVENTS"].data
event_times = lat_events["TIME"] - met
%matplotlib inline
fig, axs = plt.subplots(2, 1, sharex=True)
plt.sca(axs[0])
plt.hist(event_times)
plt.ylabel("Events")
plt.sca(axs[1])
plt.scatter(
event_times,
lat_events["ENERGY"],
marker="o",
c=lat_events["ENERGY"],
norm="log",
alpha=0.5,
zorder=20,
)
plt.yscale("log")
plt.ylabel("Energy [MeV]")
plt.xlabel("Time - T0 [s]")
plt.grid(True)
plt.show()
time -p gtmktime scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit sctable="SC_DATA" filter="(DATA_QUAL>0 || DATA_QUAL==-1) && LAT_CONFIG==1 && IN_SAA!=T && LIVETIME>0 && (ANGSEP(RA_ZENITH,DEC_ZENITH,173.136,27.7129)<=(100.0-10))" roicut=no evfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft1_tr_bn130427324_v00.fit evtable="EVENTS" outfile="gll_ft1_tr_bn130427324_v00_mkt.fit" apply_filter=yes overwrite=no header_obstimes=yes tstart=388741629.42 tstop=388742629.42 gtifile="default" chatter=2 clobber=yes debug=no gui=no mode="ql"
real 0.03
user 0.03
sys 0.00
Using 305 data
time -p gtselect infile=gll_ft1_tr_bn130427324_v00_mkt.fit outfile=gll_ft1_tr_bn130427324_v00_filt.fit ra=173.136 dec=27.7129 rad=10.0 tmin=388741629.42 tmax=388742629.42 emin=100.0 emax=100000.0 zmin=0.0 zmax=100.0 evclass=32 evtype=3 convtype=-1 phasemin=0.0 phasemax=1.0 evtable="EVENTS" chatter=2 clobber=yes debug=no gui=no mode="ql"
Done.
real 0.05
user 0.04
sys 0.00
Selected 540 events.
06:30:10 INFO Extracted 540 events download_LAT_data.py:640
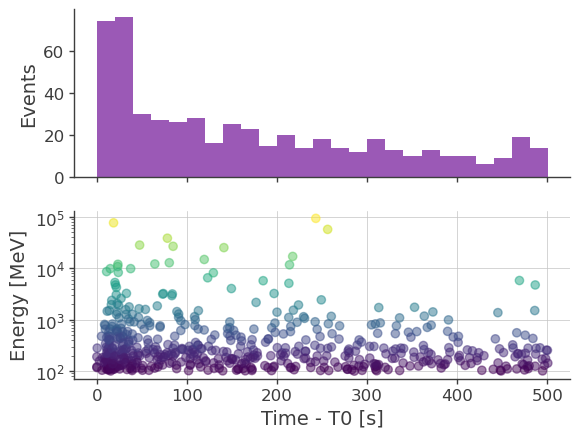
[12]:
analysis_builder = TransientLATDataBuilder(
LATdataset.grb_name,
outfile=LATdataset.grb_name,
roi=roi,
tstarts="%d" % tstart,
tstops="%d" % tstop,
irf=irf,
zmax=zmax,
galactic_model="template",
particle_model="isotr template",
datarepository=".",
)
df = analysis_builder.display(get=True)
outfile 130427324
roi 10
tstarts 0
tstops 100
zmax 100.0
emin 100.0
emax 100000.0
irf p8_transient010e
galactic_model template
particle_model isotr template
source_model PowerLaw2
tsmin 20.0
strategy time
thetamax 180.0
spectralfiles no
liketype unbinned
optimizeposition no
datarepository .
ltcube
expomap
ulphindex -2
flemin 100
flemax 10000
fgl_mode fast
filter_GTI False
likelihood_profile False
remove_fits_files False
dtype: object
[13]:
LAT_observations = analysis_builder.run(recompute_intervals=True)
LAT_plugin = LAT_observations[0].to_LATLike()
06:30:11 INFO Changing permission to lat_transient_builder.py:719 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/scripts/doTimeResolvedLike.py
INFO Changing permission to lat_transient_builder.py:719 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/gtapps_mp/gtdiffrsp_mp.py
INFO Changing permission to lat_transient_builder.py:719 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/gtapps_mp/gtexpmap_mp.py
INFO Changing permission to lat_transient_builder.py:719 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/gtapps_mp/gtltcube_mp.py
INFO Changing permission to lat_transient_builder.py:719 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/gtapps_mp/gttsmap_mp.py
INFO About to run the following command: lat_transient_builder.py:724 /home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/f ermitools/GtBurst/scripts/doTimeResolvedLike.py 130427324 --outfile '130427324' --roi 10.000000 --tstarts '0' --tstops '100' --zmax 100.000000 --emin 100.000000 --emax 100000.000000 --irf 'p8_transient010e' --galactic_model 'template' --particle_model 'isotr template' --source_model 'PowerLaw2' --tsmin 20.000000 --strategy 'time' --thetamax 180.000000 --spectralfiles 'no' --liketype 'unbinned' --optimizeposition 'no' --datarepository '.' --ltcube '' --expomap '' --ulphindex -2.000000 --flemin 100.000000 --flemax 10000.000000 --fgl_mode 'fast'
INFO You have choosen to recompute the time intervals in this folder lat_transient_builder.py:736
INFO The older entries will be moved to lat_transient_builder.py:742 tmp_e93f0822-fc46-433d-99de-702e28e8be46
**********
** 1 **SET PRINT .000
**********
**********
** 2 **SET NOWARN
**********
PARAMETER DEFINITIONS:
NO. NAME VALUE STEP SIZE LIMITS
1 'Integral ' .10000E-01 1.0000 .10000E-04 1000.0
2 'Index ' -2.0000 1.0000 -6.0000 .10000E-01
3 'Value ' 1.0000 1.0000 .70000 1.3000
4 'Normalizat' 1.0000 1.0000 .10000 10.000
**********
** 3 **SET ERR .5000
**********
**********
** 4 **SET GRAD 1.000
**********
**********
** 5 **MINIMIZE 600.0 2.000
**********
MIGRAD MINIMIZATION HAS CONVERGED.
MIGRAD WILL VERIFY CONVERGENCE AND ERROR MATRIX.
FCN= 771.6132 FROM MIGRAD STATUS=CONVERGED 73 CALLS 74 TOTAL
EDM= .65E-05 STRATEGY= 1 ERROR MATRIX ACCURATE
EXT PARAMETER STEP FIRST
NO. NAME VALUE ERROR SIZE DERIVATIVE
1 Integral .65236 .45023E-01 .10465E-02 -.98858
2 Index -1.9023 .57046E-01 .12103E-01 -.50613E-01
3 Value 1.0010 .14592 .30825 -.20057E-02
4 Normalizat 10.0000 7.4921 .45249 -.90524E-03
ERR DEF= .500
Final values:
Integral = 0.652365
Index = -1.90233
Value = 1.00099
Normalizat = 9.99999
**********
** 6 **HESSE
**********
FCN= 771.6132 FROM HESSE STATUS=OK 23 CALLS 97 TOTAL
EDM= .63E-05 STRATEGY= 1 ERROR MATRIX ACCURATE
EXT PARAMETER INTERNAL INTERNAL
NO. NAME VALUE ERROR STEP SIZE VALUE
1 Integral .65236 .44904E-01 .46267E-05 -1.5197
2 Index -1.9023 .56932E-01 .53508E-04 .37215
3 Value 1.0010 .14382 .13784E-02 .33048E-02
4 Normalizat 10.0000 7.5269 .21625E-01 1.5685
ERR DEF= .500
Minuit fit quality: 3 estimated distance: 6.33536e-06
Minuit parameter uncertainties:
1 0.0449045
2 0.0569358
3 0.149989
4 0.0186867
Requested intervals:
------------------------------------------------------
0.0 - 100.0
Data files:
-----------
eventfile /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft1_tr_bn130427324_v00.fit
ft2file /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit
rspfile /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_cspec_tr_bn130427324_v00.rsp
cspecfile /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_cspec_tr_bn130427324_v00.pha
ROI:
-----
R.A. 173.136
Dec. 27.7129
Radius 10.0
Interval # 1 (0.0-100.0):
-----------------------
-> gtdocountsmap.py rad='10.0' eventfile='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft1_tr_bn130427324_v00.fit' zmax='100.0' thetamax='180.0' emin='100.0' emax='100000.0' skymap='130427324_LAT_skymap_0.0-100.0.fit' rspfile='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_cspec_tr_bn130427324_v00.rsp' strategy='time' ft2file='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit' tstart='0.0' tstop='100.0' ra='173.136' dec='27.7129' irf='p8_transient010e' allowEmpty='no'
time -p gtmktime scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit sctable="SC_DATA" filter="(DATA_QUAL>0 || DATA_QUAL==-1) && LAT_CONFIG==1 && IN_SAA!=T && LIVETIME>0 && (ANGSEP(RA_ZENITH,DEC_ZENITH,173.136,27.7129)<=(100.0-10.0))" roicut=no evfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft1_tr_bn130427324_v00.fit evtable="EVENTS" outfile="gll_ft1_tr_bn130427324_v00_mkt.fit" apply_filter=yes overwrite=no header_obstimes=yes tstart=388741629.42 tstop=388741729.42 gtifile="default" chatter=2 clobber=yes debug=no gui=no mode="ql"
real 0.03
user 0.03
sys 0.00
Using 305 data
time -p gtselect infile=gll_ft1_tr_bn130427324_v00_mkt.fit outfile=gll_ft1_tr_bn130427324_v00_filt.fit ra=173.136 dec=27.7129 rad=10.0 tmin=388741629.42 tmax=388741729.42 emin=100.0 emax=100000.0 zmin=0.0 zmax=100.0 evclass=32 evtype=3 convtype=-1 phasemin=0.0 phasemax=1.0 evtable="EVENTS" chatter=2 clobber=yes debug=no gui=no mode="ql"
Done.
real 0.05
user 0.04
sys 0.00
Selected 233 events.
time -p gtbin evfile=gll_ft1_tr_bn130427324_v00_filt.fit scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit outfile=130427324_LAT_skymap_0.0-100.0.fit algorithm="CMAP" ebinalg="LOG" emin=30.0 emax=200000.0 ebinfile=NONE tbinalg="LIN" tbinfile=NONE nxpix=101 nypix=101 binsz=0.2 coordsys="CEL" xref=173.136 yref=27.7129 axisrot=0.0 rafield="RA" decfield="DEC" proj="AIT" hpx_ordering_scheme="RING" hpx_order=3 hpx_ebin=yes hpx_region="" evtable="EVENTS" sctable="SC_DATA" efield="ENERGY" tfield="TIME" chatter=2 clobber=yes debug=no gui=no mode="ql"
This is gtbin version HEAD
real 0.05
user 0.04
sys 0.01
Counts map saved in 130427324_LAT_skymap_0.0-100.0.fit
Total number of events in the counts map: 233
Total time in Good Time Intervals: 100.0
-> gtbuildxmlmodel xmlmodel='130427324_LAT_xmlmodel_0.0-100.0.xml' filteredeventfile='gll_ft1_tr_bn130427324_v00_filt.fit' galactic_model='template' particle_model='isotr template' ra='173.136' dec='27.7129' fgl_mode='fast' ft2file='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit' source_model='PowerLaw2'
Found Isotropic template for irf P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/iso_P8R3_TRANSIENT010E_V3_v1.txt
Found Galactic template for IRF. P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/gll_iem_v07.fits
Cutting the template around the ROI:
addFGLsources( 173.136 27.7129 18.0 130427324_LAT_xmlmodel_0.0-100.0.xml 100.0 )
Kept 0 point sources from the FGL catalog
-> gtdolike.py spectralfiles='no' xmlmodel='130427324_LAT_xmlmodel_0.0-100.0.xml' liketype='unbinned' filteredeventfile='gll_ft1_tr_bn130427324_v00_filt.fit' rspfile='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_cspec_tr_bn130427324_v00.rsp' showmodelimage='no' tsmin='20.0' optimizeposition='no' ft2file='/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit' skymap='130427324_LAT_skymap_0.0-100.0.fit' flemin='100.000000' flemax='10000.000000'
time -p gtltcube evfile="/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit" evtable="EVENTS" scfile=__ft2temp.fits sctable="SC_DATA" outfile=gll_ft1_tr_bn130427324_v00_filt_ltcube.fit dcostheta=0.025 binsz=1.0 phibins=1 tmin=0.0 tmax=0.0 file_version="1" zmin=0.0 zmax=180.0 chatter=2 clobber=yes debug=no gui=no mode="ql"
Working on file __ft2temp.fits
......!
real 0.34
user 0.17
sys 0.17
/home/runner/miniconda3/envs/test_env/lib/python3.11/site-packages/fermitools/GtBurst/gtapps_mp/gtexpmap_mp.py 40 40 2 2 /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit gll_ft1_tr_bn130427324_v00_filt_ltcube.fit P8R3_TRANSIENT010E_V3 20.0 20 gll_ft1_tr_bn130427324_v00_filt_expomap.fit
Starting calculation of region 20.0,20.0 to 40.0,40.0
Completed calculation of region 20.0,20.0 to 40.0,40.0
Starting calculation of region 0.0,0.0 to 20.0,20.0
Completed calculation of region 0.0,0.0 to 20.0,20.0
Starting calculation of region 0.0,20.0 to 20.0,40.0
Completed calculation of region 0.0,20.0 to 20.0,40.0
Starting calculation of region 20.0,0.0 to 40.0,20.0
Completed calculation of region 20.0,0.0 to 40.0,20.0
Spawning 4 jobs...
Combining temporary files...
Deleting temporary files...
time -p gtdiffrsp evfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit evtable="EVENTS" scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit sctable="SC_DATA" srcmdl=130427324_LAT_xmlmodel_0.0-100.0.xml irfs="P8R3_TRANSIENT010E_V3" evclsmin="INDEF" evclass="INDEF" evtype="INDEF" convert=no chatter=2 clobber=yes debug=no gui=no mode="ql"
adding source GalacticTemplate
adding source IsotropicTemplate
Working on...
/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit......................!
real 7.70
user 7.37
sys 0.33
Loading python Likelihood interface...
Applying a Gaussian prior with sigma 0.15 on the normalization of the Galactic Template
{'Norm': 1.0, 'Mean': 1.0, 'Sigma': 0.15, 'Offset': 0.0}
Likelihood settings:
Event file(s): /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit
Spacecraft file(s): /home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit
Exposure map: gll_ft1_tr_bn130427324_v00_filt_expomap.fit
Exposure cube: gll_ft1_tr_bn130427324_v00_filt_ltcube.fit
IRFs: P8R3_TRANSIENT010E_V3
Source model file: 130427324_LAT_xmlmodel_0.0-100.0.xml
Optimizer: Minuit
Performing likelihood fit...
Data 233.0
srcName 228.0711654565354
srcName 0.21216428870537518
srcName 3.8519232874938027
Resid 233.0 232.13525303273457 5.00806149863562
plotting GRB spectrum
srcName 228.0711654565354
|--------------------|---------------|----------| ----------|----------|------|
| Source name| Par. Name| Value| Error| Units| TS|
|--------------------|---------------|----------|------------|----------|------|
|GRB | | | | | 2540|
| | Integral| 0.000652| 4.49e-05| ph./cm2/s| |
| | Index| -1.9| 0.0569| -| |
| | LowerLimit| 100|n.a. (fixed)| MeV| |
| | UpperLimit| 1e+05|n.a. (fixed)| MeV| |
| | Energy flux| 5.5e-07| 4.99e-08| erg/cm2/s| |
| | Photon flux| 0.000643| 4.49e-05| ph./cm2/s| |
|GalacticTemplate | | | | | 1|
| | Value| 1| 0.15| -| |
| | Energy flux| 3.76e-07| 5.64e-08| erg/cm2/s| |
| | Photon flux| 0.00052| 7.79e-05| ph./cm2/s| |
|IsotropicTemplate | | | | | 4|
| | Normalization| 10| 0.0187| -| |
| | Energy flux| 7.82e-07| 1.46e-09| erg/cm2/s| |
| | Photon flux| 0.00158| 2.96e-06| ph./cm2/s| |
--------------------------------------------------------------------------------
*** All fluxes and upper limits have been computed in the 100.0 - 10000.0 energy range.
*** Upper limits (if any) are computed assuming a photon index of -2.0, with the 95.0 % c.l.
Log(likelihood) = 771.6131616797278
time -p gtsrcprob evfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt.fit evtable="EVENTS" scfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/bn130427324/gll_ft2_tr_bn130427324_v00.fit sctable="SC_DATA" outfile=/home/runner/work/threeML/threeML/docs/md_docs/slow_execute/interval0.0-100.0/gll_ft1_tr_bn130427324_v00_filt_prob.fit srcmdl=gll_ft1_tr_bn130427324_v00_filt_likeRes.xml irfs="CALDB" evtype="INDEF" srclist= chatter=2 clobber=yes debug=no gui=no mode="ql"
real 3.65
user 3.28
sys 0.37
06:30:41 INFO The ft2 file does not exist. Please examine! lat_transient_builder.py:818
INFO we will grab the data file for you. lat_transient_builder.py:819
INFO copied ./bn130427324/gll_ft2_tr_bn130427324_v00.fit to lat_transient_builder.py:838 interval0.0-100.0/gll_ft2_tr_bn130427324_v00.fit
FermiLATLike - GTI SUM = 100.0
Define the first model to test
[14]:
band = Band()
band.alpha.prior = Truncated_gaussian(lower_bound=-1.5, upper_bound=1, mu=-1, sigma=0.5)
band.beta.prior = Truncated_gaussian(lower_bound=-5, upper_bound=-1.6, mu=-2, sigma=0.5)
band.xp.prior = Log_normal(mu=2, sigma=1)
band.xp.bounds = (None, None)
band.K.prior = Log_uniform_prior(lower_bound=1e-10, upper_bound=1e3)
source = PointSource(trigger_name, ra, dec, spectral_shape=band)
band_model = Model(source)
06:30:41 WARNING We have set the min_value of Band.xp to 1e-99 because there was a postive parameter.py:715 transform
WARNING We have set the min_value of Band.xp to 1e-99 because there was a postive parameter.py:715 transform
Run the joint fit
[15]:
datalist_1 = DataList(*gbm_plugins, lle_plugin, LAT_plugin)
jl_1 = JointLikelihood(band_model, datalist_1, verbose=False)
band_model.display()
print(jl_1.data_list.keys())
# This is needed to fix the galactic template to 1 (if needed)
band_model[LAT_plugin.get_name() + "_GalacticTemplate_Value"].value = 1.0
band_model[LAT_plugin.get_name() + "_GalacticTemplate_Value"].fix = True
band_model[LAT_plugin.get_name() + "_IsotropicTemplate_Normalization"].fix = False
# sbpl_model.display(complete=True)
jl_1.set_minimizer("minuit")
jl_1.fit()
Found Isotropic template for irf P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/iso_P8R3_TRANSIENT010E_V3_v1.txt
Found Galactic template for IRF. P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/gll_iem_v07.fits
Cutting the template around the ROI:
06:30:45 INFO set the minimizer to minuit joint_likelihood.py:1017
Model summary:
Free parameters (7):
Fixed parameters (6):
(abridged. Use complete=True to see all fixed parameters)
Properties (0):
(none)
Linked parameters (0):
(none)
Independent variables:
(none)
Linked functions (0):
(none)
| N | |
|---|---|
| Point sources | 1 |
| Extended sources | 0 |
| Particle sources | 0 |
Free parameters (7):
| value | min_value | max_value | unit | |
|---|---|---|---|---|
| bn130427324.spectrum.main.Band.K | 0.0001 | 0.0 | None | keV-1 s-1 cm-2 |
| bn130427324.spectrum.main.Band.alpha | -1.0 | -1.5 | 3.0 | |
| bn130427324.spectrum.main.Band.xp | 500.0 | 0.0 | None | keV |
| bn130427324.spectrum.main.Band.beta | -2.0 | -5.0 | -1.6 | |
| cons_lat_lle | 1.0 | 0.8 | 1.2 | |
| LAT0X100_GalacticTemplate_Value | 1.0 | 0.5 | 1.5 | |
| LAT0X100_IsotropicTemplate_Normalization | 1.0 | 0.1 | 5.0 |
Fixed parameters (6):
(abridged. Use complete=True to see all fixed parameters)
Properties (0):
(none)
Linked parameters (0):
(none)
Independent variables:
(none)
Linked functions (0):
(none)
odict_keys(['n6', 'n9', 'b1', 'lat_lle', 'LAT0X100'])
INFO set the minimizer to MINUIT joint_likelihood.py:1034
06:30:46 WARNING 74.56 percent of samples have been thrown away because they failed the analysis_results.py:1645 constraints on the parameters. This results might not be suitable for error propagation. Enlarge the boundaries until you loose less than 1 percent of the samples.
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| bn130427324.spectrum.main.Band.K | (1.496 +/- 0.004) x 10^-1 | 1 / (keV s cm2) |
| bn130427324.spectrum.main.Band.alpha | (-9.12 +/- 0.05) x 10^-1 | |
| bn130427324.spectrum.main.Band.xp | (7.76 +/- 0.07) x 10^2 | keV |
| bn130427324.spectrum.main.Band.beta | -2.837 +/- 0.011 | |
| cons_lat_lle | (8.0000 +/- 0.0005) x 10^-1 | |
| LAT0X100_IsotropicTemplate_Normalization | 5.000 +/- 0.018 |
Correlation matrix:
| 1.00 | 0.55 | -0.84 | 0.32 | -0.00 | -0.00 |
| 0.55 | 1.00 | -0.78 | 0.26 | -0.00 | -0.00 |
| -0.84 | -0.78 | 1.00 | -0.41 | 0.00 | 0.00 |
| 0.32 | 0.26 | -0.41 | 1.00 | -0.00 | -0.00 |
| -0.00 | -0.00 | 0.00 | -0.00 | 1.00 | 0.00 |
| -0.00 | -0.00 | 0.00 | -0.00 | 0.00 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| n6 | 1109.980692 |
| n9 | 1063.105290 |
| b1 | 896.397958 |
| lat_lle | 85.188113 |
| LAT0X100 | 1913.535513 |
| total | 5068.207566 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 10148.573622 |
| BIC | 10174.131120 |
[15]:
( value negative_error \
bn130427324.spectrum.main.Band.K 0.149628 -0.000386
bn130427324.spectrum.main.Band.alpha -0.912123 -0.004352
bn130427324.spectrum.main.Band.xp 775.990259 -6.369709
bn130427324.spectrum.main.Band.beta -2.837093 -0.010239
cons_lat_lle 0.800000 0.000009
LAT0X100_IsotropicTemplate_Normalization 4.999963 -0.024964
positive_error error \
bn130427324.spectrum.main.Band.K 0.000376 0.000381
bn130427324.spectrum.main.Band.alpha 0.004523 0.004438
bn130427324.spectrum.main.Band.xp 6.375007 6.372358
bn130427324.spectrum.main.Band.beta 0.009954 0.010096
cons_lat_lle 0.000068 0.000038
LAT0X100_IsotropicTemplate_Normalization -0.003380 0.010792
unit
bn130427324.spectrum.main.Band.K 1 / (keV s cm2)
bn130427324.spectrum.main.Band.alpha
bn130427324.spectrum.main.Band.xp keV
bn130427324.spectrum.main.Band.beta
cons_lat_lle
LAT0X100_IsotropicTemplate_Normalization ,
-log(likelihood)
n6 1109.980692
n9 1063.105290
b1 896.397958
lat_lle 85.188113
LAT0X100 1913.535513
total 5068.207566)
[16]:
display_spectrum_model_counts(jl_1)
[16]:
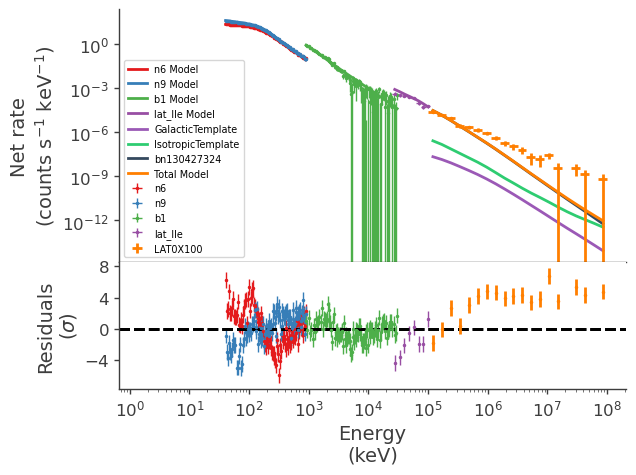
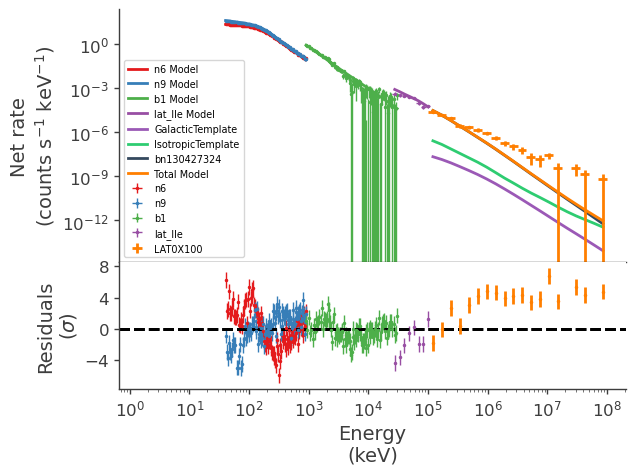
Define a second model to test
[17]:
spectrum = Band() + Powerlaw()
# spectrum.K_1 = 0.1
# spectrum.break_energy_1 = 800
# spectrum.beta_1 = -3
# spectrum.K_2 = 3
source_2 = PointSource("%s_2" % trigger_name, ra, dec, spectral_shape=spectrum)
comp_model = Model(source_2)
comp_model.display()
Model summary:
Free parameters (6):
Fixed parameters (4):
(abridged. Use complete=True to see all fixed parameters)
Properties (0):
(none)
Linked parameters (0):
(none)
Independent variables:
(none)
Linked functions (0):
(none)
| N | |
|---|---|
| Point sources | 1 |
| Extended sources | 0 |
| Particle sources | 0 |
Free parameters (6):
| value | min_value | max_value | unit | |
|---|---|---|---|---|
| bn130427324_2.spectrum.main.composite.K_1 | 0.0001 | 0.0 | None | keV-1 s-1 cm-2 |
| bn130427324_2.spectrum.main.composite.alpha_1 | -1.0 | -1.5 | 3.0 | |
| bn130427324_2.spectrum.main.composite.xp_1 | 500.0 | 10.0 | None | keV |
| bn130427324_2.spectrum.main.composite.beta_1 | -2.0 | -5.0 | -1.6 | |
| bn130427324_2.spectrum.main.composite.K_2 | 1.0 | 0.0 | 1000.0 | keV-1 s-1 cm-2 |
| bn130427324_2.spectrum.main.composite.index_2 | -2.01 | -10.0 | 10.0 |
Fixed parameters (4):
(abridged. Use complete=True to see all fixed parameters)
Properties (0):
(none)
Linked parameters (0):
(none)
Independent variables:
(none)
Linked functions (0):
(none)
Run the joint fit again
[18]:
datalist_2 = DataList(*gbm_plugins, lle_plugin, LAT_plugin)
jl_2 = JointLikelihood(comp_model, datalist_2, verbose=False)
comp_model[LAT_plugin.get_name() + "_GalacticTemplate_Value"].value = 1.0
comp_model[LAT_plugin.get_name() + "_GalacticTemplate_Value"].fix = True
comp_model[LAT_plugin.get_name() + "_IsotropicTemplate_Normalization"].fix = False
jl_2.set_minimizer("minuit")
jl_2.fit()
Found Isotropic template for irf P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/iso_P8R3_TRANSIENT010E_V3_v1.txt
Found Galactic template for IRF. P8R3_TRANSIENT010E_V3: /home/runner/miniconda3/envs/test_env/share/fermitools/refdata/fermi/galdiffuse/gll_iem_v07.fits
Cutting the template around the ROI:
06:30:52 INFO set the minimizer to minuit joint_likelihood.py:1017
INFO set the minimizer to MINUIT joint_likelihood.py:1034
06:30:53 WARNING 74.8 percent of samples have been thrown away because they failed the analysis_results.py:1645 constraints on the parameters. This results might not be suitable for error propagation. Enlarge the boundaries until you loose less than 1 percent of the samples.
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| bn130427324_2.spectrum.main.composite.K_1 | (1.482 +/- 0.005) x 10^-1 | 1 / (keV s cm2) |
| bn130427324_2.spectrum.main.composite.alpha_1 | (-9.19 +/- 0.05) x 10^-1 | |
| bn130427324_2.spectrum.main.composite.xp_1 | (7.96 +/- 0.07) x 10^2 | keV |
| bn130427324_2.spectrum.main.composite.beta_1 | -3.022 +/- 0.030 | |
| bn130427324_2.spectrum.main.composite.K_2 | 1.1 -0.6 +1.5 | 1 / (keV s cm2) |
| bn130427324_2.spectrum.main.composite.index_2 | -1.71 +/- 0.06 | |
| cons_lat_lle | (8.000 +/- 0.034) x 10^-1 | |
| LAT0X100_IsotropicTemplate_Normalization | 5.000 +/- 0.008 |
Correlation matrix:
| 1.00 | 0.26 | -0.72 | 0.56 | -0.61 | 0.59 | 0.00 | -0.00 |
| 0.26 | 1.00 | -0.73 | 0.06 | 0.25 | -0.26 | 0.00 | 0.00 |
| -0.72 | -0.73 | 1.00 | -0.38 | 0.10 | -0.08 | -0.00 | -0.00 |
| 0.56 | 0.06 | -0.38 | 1.00 | -0.57 | 0.52 | 0.01 | -0.00 |
| -0.61 | 0.25 | 0.10 | -0.57 | 1.00 | -0.99 | -0.00 | 0.00 |
| 0.59 | -0.26 | -0.08 | 0.52 | -0.99 | 1.00 | 0.00 | -0.00 |
| 0.00 | 0.00 | -0.00 | 0.01 | -0.00 | 0.00 | 1.00 | 0.00 |
| -0.00 | 0.00 | -0.00 | -0.00 | 0.00 | -0.00 | 0.00 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| n6 | 1114.367049 |
| n9 | 1061.440251 |
| b1 | 880.522911 |
| lat_lle | 70.125802 |
| LAT0X100 | 1811.599469 |
| total | 4938.055482 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 9892.383691 |
| BIC | 9926.398949 |
[18]:
( value negative_error \
bn130427324_2.spectrum.main.composite.K_1 0.148244 -0.000501
bn130427324_2.spectrum.main.composite.alpha_1 -0.918506 -0.004899
bn130427324_2.spectrum.main.composite.xp_1 796.175741 -6.847524
bn130427324_2.spectrum.main.composite.beta_1 -3.021983 -0.032184
bn130427324_2.spectrum.main.composite.K_2 1.105333 -0.608719
bn130427324_2.spectrum.main.composite.index_2 -1.707909 -0.059393
cons_lat_lle 0.800015 0.000696
LAT0X100_IsotropicTemplate_Normalization 4.999949 -0.011301
positive_error error \
bn130427324_2.spectrum.main.composite.K_1 0.000502 0.000501
bn130427324_2.spectrum.main.composite.alpha_1 0.004691 0.004795
bn130427324_2.spectrum.main.composite.xp_1 7.190828 7.019176
bn130427324_2.spectrum.main.composite.beta_1 0.029176 0.030680
bn130427324_2.spectrum.main.composite.K_2 1.439043 1.023881
bn130427324_2.spectrum.main.composite.index_2 0.056554 0.057974
cons_lat_lle 0.004650 0.002673
LAT0X100_IsotropicTemplate_Normalization -0.001310 0.004996
unit
bn130427324_2.spectrum.main.composite.K_1 1 / (keV s cm2)
bn130427324_2.spectrum.main.composite.alpha_1
bn130427324_2.spectrum.main.composite.xp_1 keV
bn130427324_2.spectrum.main.composite.beta_1
bn130427324_2.spectrum.main.composite.K_2 1 / (keV s cm2)
bn130427324_2.spectrum.main.composite.index_2
cons_lat_lle
LAT0X100_IsotropicTemplate_Normalization ,
-log(likelihood)
n6 1114.367049
n9 1061.440251
b1 880.522911
lat_lle 70.125802
LAT0X100 1811.599469
total 4938.055482)
[19]:
display_spectrum_model_counts(jl_2, show_residuals=True)
[19]:
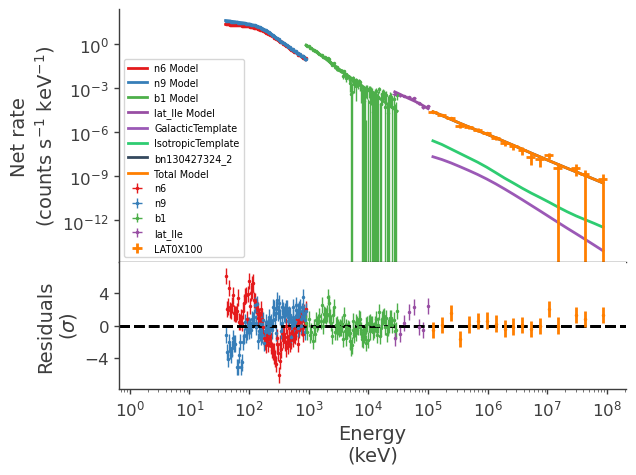
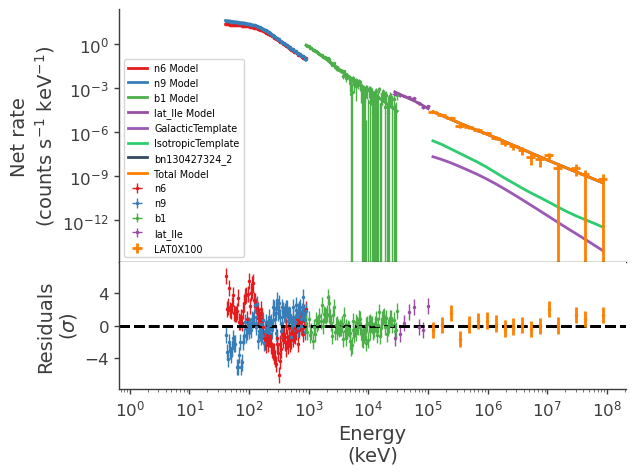
[20]:
jl_1.results.get_statistic_measure_frame()
[20]:
| statistical measures | |
|---|---|
| AIC | 10148.573622 |
| BIC | 10174.131120 |
[21]:
jl_2.results.get_statistic_measure_frame()
[21]:
| statistical measures | |
|---|---|
| AIC | 9892.383691 |
| BIC | 9926.398949 |
Plot the two components of the best-fit spectrum
[22]:
fig = plot_spectra(
jl_2.results,
ene_min=10 * u.keV,
ene_max=100 * u.GeV,
flux_unit="erg2/(cm2 s keV)",
fit_cmap="viridis",
contour_cmap="viridis",
contour_style_kwargs=dict(alpha=0.1),
use_components=True,
components_to_use=["total", "Band", "Powerlaw"],
)
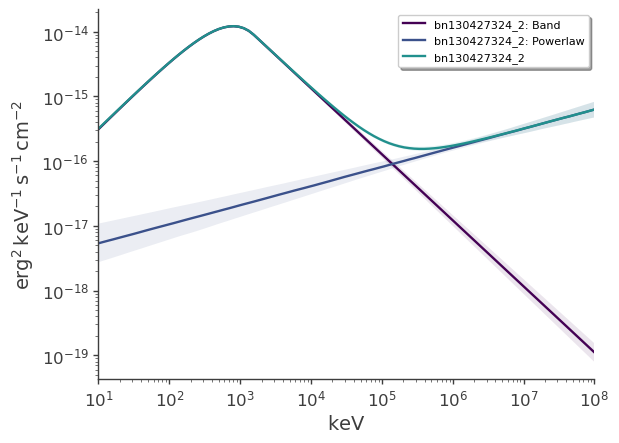
Compute the flux
[23]:
jl_2.results.get_flux(
ene_min=10 * u.keV, ene_max=100 * u.GeV, use_components=True, flux_unit="1/(cm2 s)"
)
[23]:
| flux | low bound | hi bound | |
|---|---|---|---|
| bn130427324_2: Band | 56.28494352509192 1 / (s cm2) | 56.101015983722455 1 / (s cm2) | 56.48717963990927 1 / (s cm2) |
| bn130427324_2: Powerlaw | 0.32109622900410895 1 / (s cm2) | 0.17608720410206355 1 / (s cm2) | 0.5946621346485931 1 / (s cm2) |
[24]:
jl_2.results.get_flux(
ene_min=10 * u.keV, ene_max=100 * u.GeV, use_components=False, flux_unit="1/(cm2 s)"
)
[24]:
| flux | low bound | hi bound | |
|---|---|---|---|
| bn130427324_2: total | 56.66211916676208 1 / (s cm2) | 56.49478601414048 1 / (s cm2) | 56.86105765980553 1 / (s cm2) |